DynaMut2: Assessing changes in stability and flexibility upon single and multiple point missense mutations - Rodrigues - 2021 - Protein Science - Wiley Online Library
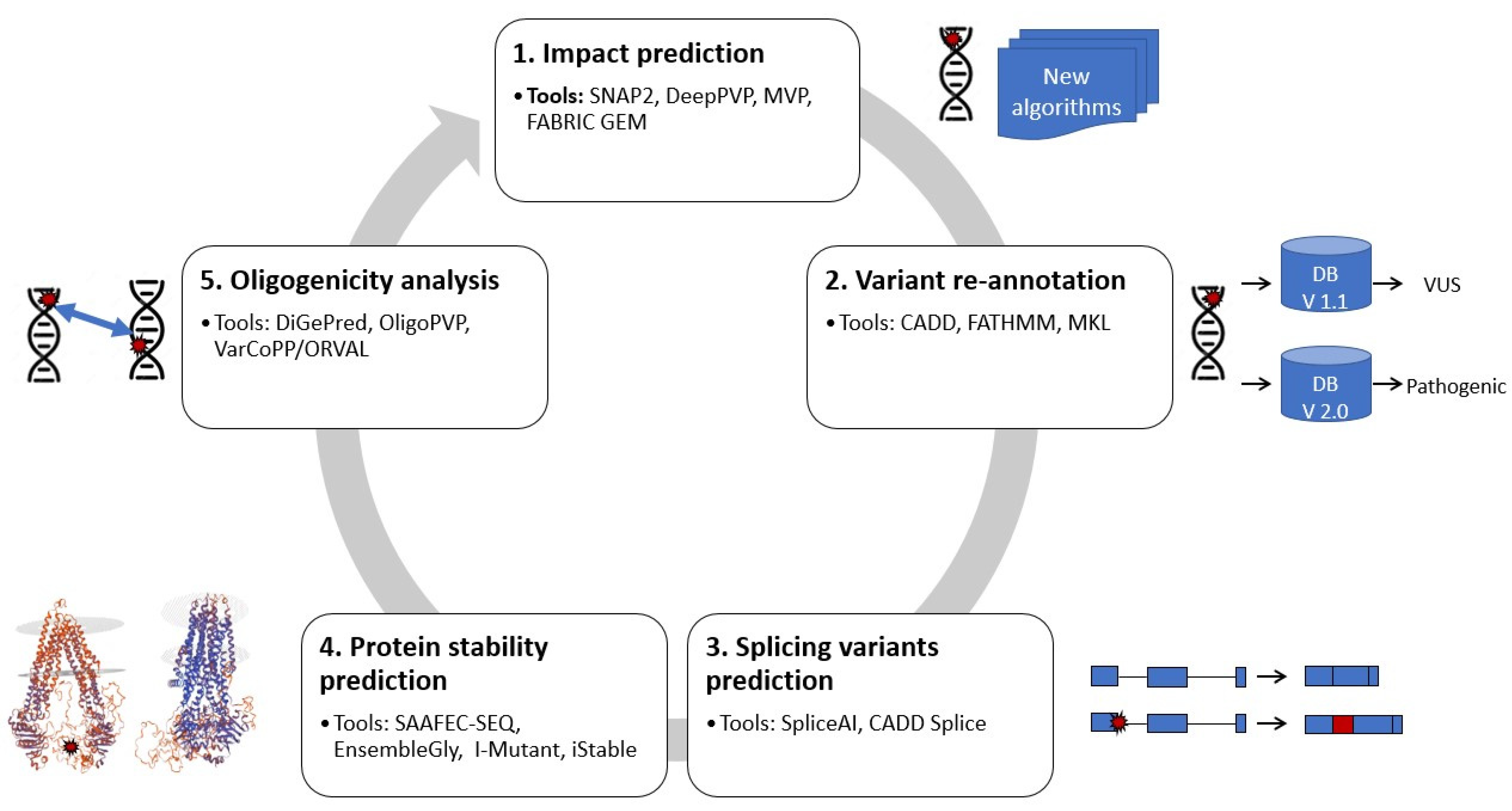
IJMS | Free Full-Text | New Developments and Possibilities in Reanalysis and Reinterpretation of Whole Exome Sequencing Datasets for Unsolved Rare Diseases Using Machine Learning Approaches
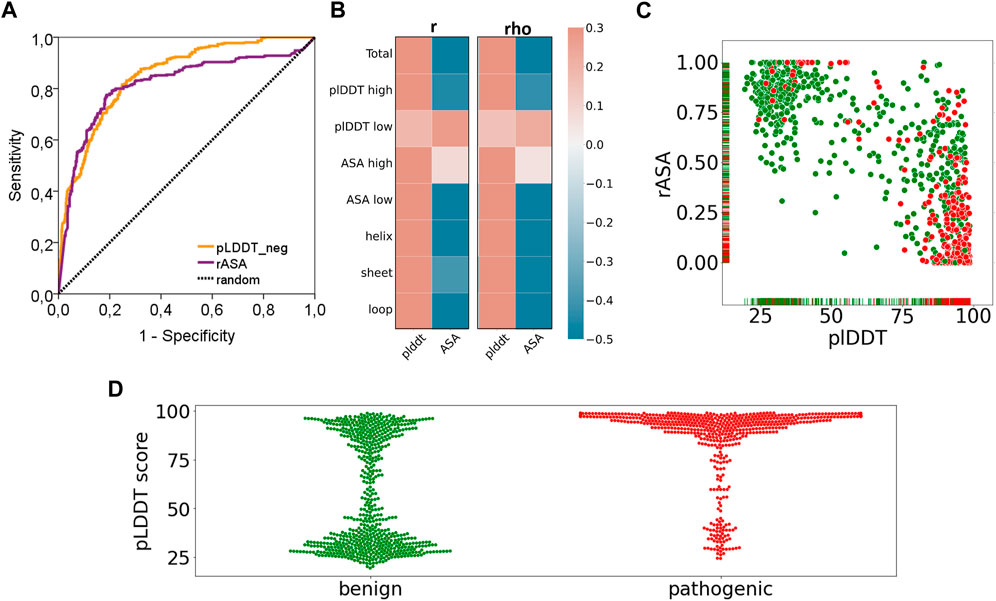
Frontiers | Evaluation of AlphaFold structure-based protein stability prediction on missense variations in cancer

Dissecting the stability determinants of a challenging de novo protein fold using massively parallel design and experimentation | PNAS
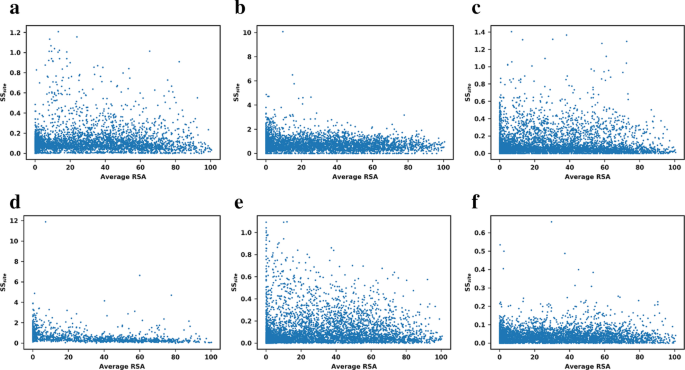
A base measure of precision for protein stability predictors: structural sensitivity | BMC Bioinformatics | Full Text

Computational Modeling of Protein Stability: Quantitative Analysis Reveals Solutions to Pervasive Problems - ScienceDirect

SCONES: Self-Consistent Neural Network for Protein Stability Prediction Upon Mutation | The Journal of Physical Chemistry B

Finding the ΔΔG spot: Are predictors of binding affinity changes upon mutations in protein–protein interactions ready for it? - Geng - 2019 - WIREs Computational Molecular Science - Wiley Online Library

Artificial intelligence challenges for predicting the impact of mutations on protein stability - ScienceDirect
Predicting changes in protein thermodynamic stability upon point mutation with deep 3D convolutional neural networks | PLOS Computational Biology

Prediction of protein stability changes upon single-point variant using 3D structure profile - ScienceDirect

Novel Algorithms and Tools for Computational Protein Design with Applications to Drug Resistance Prediction, Antibody Design, Peptide Inhibitor Design, and Protein Stability Prediction

Limitations and challenges in protein stability prediction upon genome variations: towards future applications in precision medicine - ScienceDirect

MPTherm-pred: Analysis and Prediction of Thermal Stability Changes upon Mutations in Transmembrane Proteins - ScienceDirect
Computational Modeling of Protein Stability: Quantitative Analysis Reveals Solutions to Pervasive Problems

In Silico Tools and Approaches for the Prediction of Functional and Structural Effects of Single-Nucleotide Polymorphisms on Proteins: An Expert Review | OMICS: A Journal of Integrative Biology
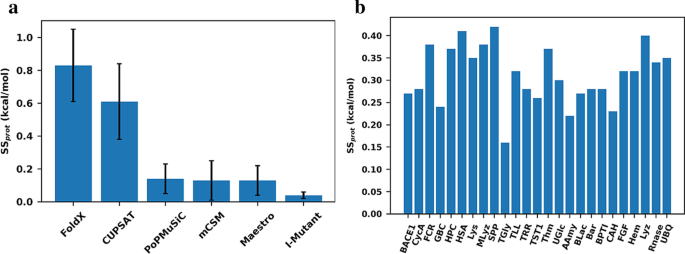
A base measure of precision for protein stability predictors: structural sensitivity | BMC Bioinformatics | Full Text

Predicting and interpreting large-scale mutagenesis data using analyses of protein stability and conservation - ScienceDirect
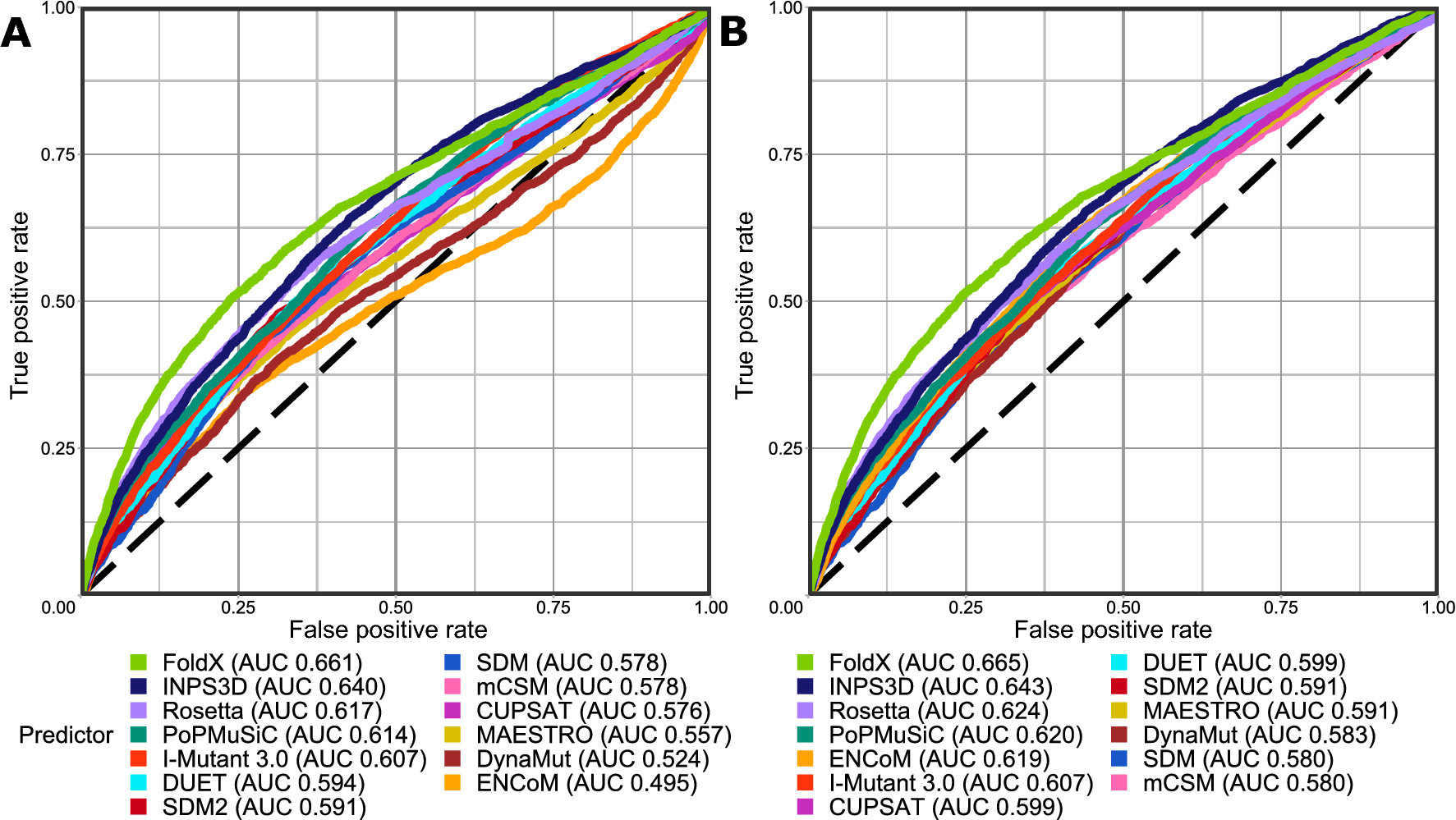
Identification of pathogenic missense mutations using protein stability predictors | Scientific Reports